Fluorescent proteins or compounds getting in the way of DSF? Try Stargazer.
Stargazer-2's non-specific, label-free technology is unaffected by fluorescent samples. Contact us and discover the benefits of adding Stargazer to your lab bench.
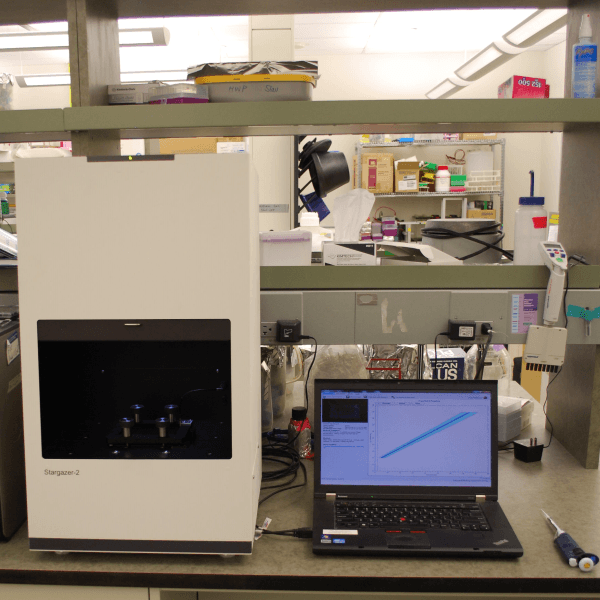


Next-generation differential static light scattering
We took the original Stargazer-384 instrument and redesigned the hardware and software from scratch. The result is Stargazer-2 — faster, more accurate, and easier to use.
Advanced Software
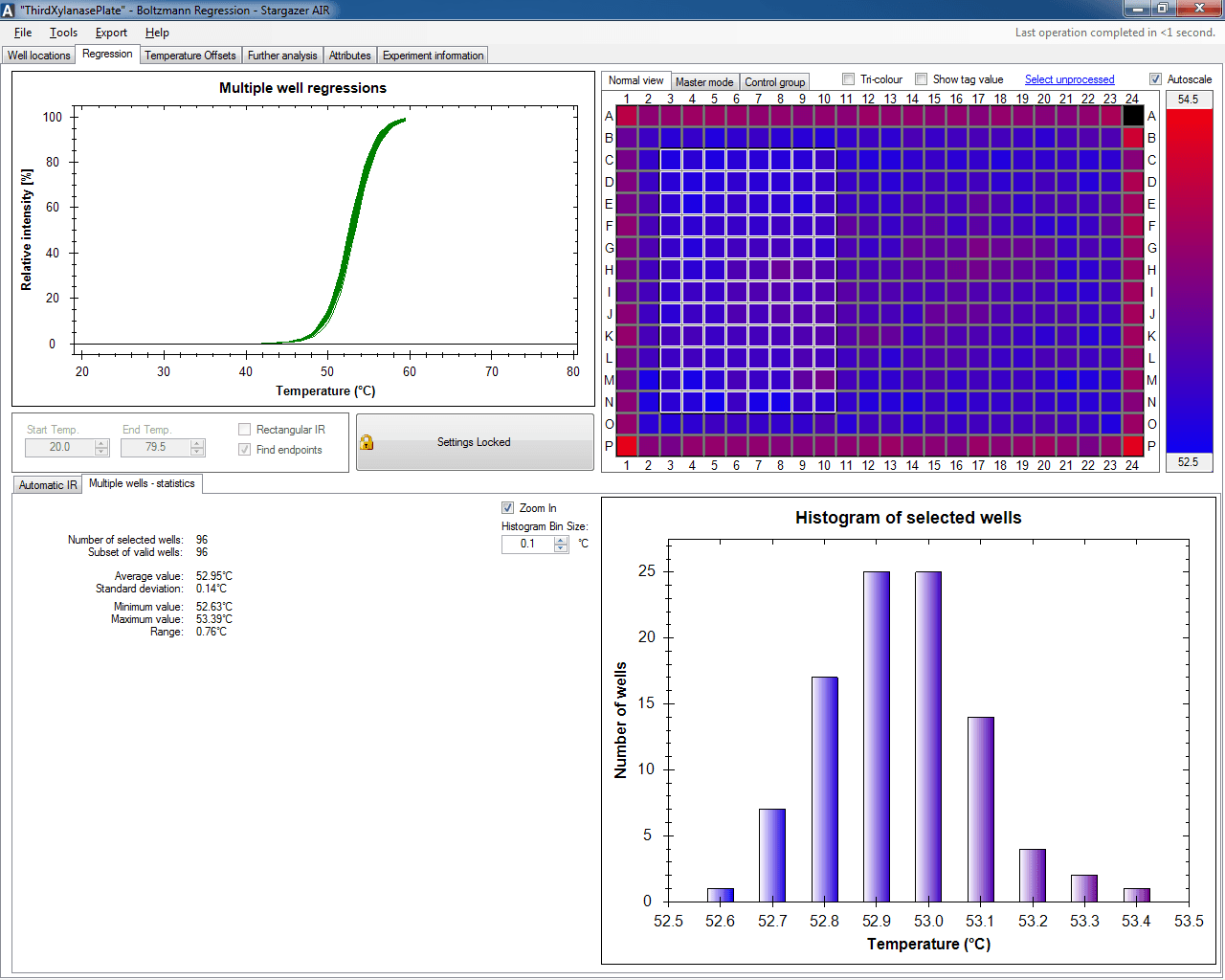
Stargazer's Magellanic control software allows users to customize the operation of Stargazer-2 to their needs with method-based experiments.
Stargazer-AIR automatically processes experiment results and provides the user with an intuitive graphical interface to review their results.
A modular plugin architecture allows users to export results to Excel and BioActive. LIMS integration is possible with custom plugins.
Online software updates ensure the best user experience.